I2M team
Molecular and metabolic engineering
A team dedicated to metabolic and protein engineering
Molecular and metabolic engineering
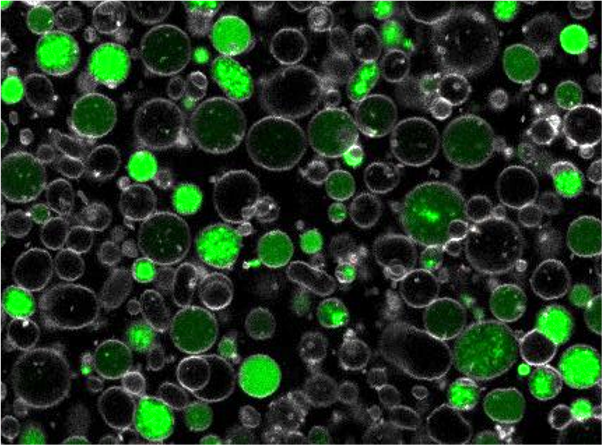
Our main lines of research are aimed at the design and construction of optimized biological systems, whether macromolecules (molecular engineering) or microorganisms (metabolic engineering). We rely on our expertise in protein engineering, in particular cytochrome P450 and bacterial microcompartment shell proteins, and in synthetic biology, in particular on the carotenoid and terpene production pathway in the yeast Saccharomyces cerevisiae. We analyze how enzymatic and cellular parameters are responsible for the efficiency of a synthetic metabolic pathway in order to reduce its physiological and metabolic impact on the host cell. To do so, we seek to control enzyme activities within synthetic pathways (quantity, activity, promiscuity, assembly) or optimize yeast strains (expression regulation, subcellular localization, etc.).
- ANR
- TWB
- Sanofi
- Servier
- Adisseo
- Rules for assembly of bacterial protein microcompartments.
- Optimization of the carotene pathway in the yeast Saccharomyces cerevisiae.
- Effect of intracellular localization in the host organism of products of heterologous metabolism
- Biosensors and metabolic engineering in yeast.
- Study of different linkers to artificially associate enzymes involved in carotene biosynthesis
- Study of multiple mutations of the active site of a terpene synthetase on its metabolic profile. In silico study of the assembly modalities of protein complexes using AlphaFold2, a deep-learning artificial intelligence tool.
- Lucie Barthe (sur place): Study of the spontaneous assembly of individual bacterial micro-compartment shell components.
- Aurélie Bouin (à Singapour): Amélioration du flux métabolique de la voie des caroténoïdes.