Les écosystèmes microbiens représentent une mine d’or pour la prospection de nouvelles fonctions d’intérêt biotechnologique. Pourtant, la majorité des espèces qui composent ces communautés complexes ne sont pas cultivées et constituent une boîte noire dont le fonctionnement est difficile, voire impossible à étudier. Pour faire face à ces enjeux, le TBI et TWB , en collaboration avec l’unité INSERM I2MC, ont intégré microbiomics et microfluidique en gouttes pour développer une nouvelle stratégie de criblage à ultra-haut débit pour la culturomique, la métagenomique fonctionnelle, l’ingénierie d’enzymes, de souches et de consortia microbiens. Deux technologies de rupture, qui ne reposent ni sur l’utilisation de substrats fluorogéniques ni sur des banques fluorescentes, ont été développées et brevetées par l’INRAE. Elles sont actuellement exploitées dans le cadre du projet ANR CAZIBD pour décrypter le dialogue hôte-microbiote-aliment, et du projet européen BLUETOOLS dédié à l’étude et l’exploitation biotechnologique des écosystèmes marins.
Microfluidics allows coupling ultra-high throughput isolation and culture of microorganisms in drops with phenotypic, taxonomic and genomic characterization. However, existing technologies require either labeled recombinant cell banks, cell survival markers, or fluorogenic or chromogenic probes or substrates. They are therefore not adapted to the culture and screening of native microorganisms, nor to the use of non-chemically modified substrates naturally present in microbial ecosystems. These barriers limit the understanding of the functioning of ecosystems to their biotechnological exploitation.
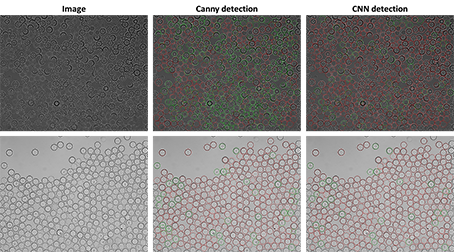
Two revolutionary technologies, based on different principles, have been developed. The first integrates drop microfluidics, imaging, artificial intelligence and flow cytometry for automated detection and sorting of positive drops, at a rate of about 105 drops per hour. The second technology, dedicated to culturomics and strain and consortia engineering, directly couples drop microfluidics and flow cytometry for the selection of growing microorganisms, at a rate of 106 drops per hour. With a picoliter scale, a million times smaller than the sample volumes used with conventional phenotypic screening techniques, these technologies are compatible with any type of substrate at any cost, and the screening of any phenotype, as long as it can be detected by confocal microscopy or linked to microbial cell growth.
Les technologies développées peuvent être exploitées pour la métagénomique fonctionnelle, la culturomique, l’ingénierie d’enzymes même en utilisant des systèmes acellulaires, l’ingénierie de souches ou de consortia, par exemple pour la production d’antimicrobiens ou pour la dégradation de polymères synthétiques polluants tels que les plastiques.
Valorization
Exploitation of these technologies in the framework of the ANR CAZIBD (Mucus-degrading enzymes and their role in intestinal inflammation) and the HORIZON BLUETOOLS project (Innovative tools for sustainable exploration of marine microbiominnovative tools for sustainable exploration of marine microbiomes : towards a circular blue bioeconomy and healthier marine environments)
Patents
- S. Lajus, Flores-Flores R., Lestrade D., Deroite A., Dagkesamanskaya, Potocki-Veronese G. A
new droplet micro-to-milli-fluidics-based process to screen for phenotypes or biological processes.
Brevet européen déposé par INRAE le 20/05/2022 sous le N°EP22305758.9
- S. Lajus, Lestrade D., Deroite A., Dagkesamanskaya, Potocki-Veronese G. New milli-to-microfluidics-based process to screen microbial growth in droplets. Brevet européen déposé par INRAE le 20/05/2022 sous le N° EP22305759.7
Contacts
Gabrielle Potocki-Veronese (veronese@insa-toulouse.fr)
Sophie Lajus (lajus@insa-toulouse.fr)
Delphine Lestrade (delphine.lestrade@inrae.fr)
Légende et copyright image : Apprentissage pour la détection automatisée et le tri de gouttes pour le criblage à ultra-haut débit de fonctions microbiennes (Sophie Lajus et Remy Flores-Flores)